





Acta Veterinaria et Zootechnica Sinica ›› 2025, Vol. 56 ›› Issue (8): 3561-3577.doi: 10.11843/j.issn.0366-6964.2025.08.001
• Review • Previous Articles Next Articles
LIU Can1,2,3( ), SU Yixin1,2,3, JING Xianjin1,2,3, LI Wenze1,2,3, YANG Lepu1,2,3, WANG Ruijun1,2,3, ZHANG Yanjun1,2,3, WANG Zhiying1,2,3, LÜ Qi1,2,3, SU Rui1,2,3,*(
), SU Yixin1,2,3, JING Xianjin1,2,3, LI Wenze1,2,3, YANG Lepu1,2,3, WANG Ruijun1,2,3, ZHANG Yanjun1,2,3, WANG Zhiying1,2,3, LÜ Qi1,2,3, SU Rui1,2,3,*( )
)
Received:2024-12-13
Online:2025-08-23
Published:2025-08-28
Contact:
SU Rui
E-mail:15847756571@163.com;suruiyu@126.com
CLC Number:
LIU Can, SU Yixin, JING Xianjin, LI Wenze, YANG Lepu, WANG Ruijun, ZHANG Yanjun, WANG Zhiying, LÜ Qi, SU Rui. Research Progress of Epigenetics in Sheep and Goats Genetic Breeding[J]. Acta Veterinaria et Zootechnica Sinica, 2025, 56(8): 3561-3577.
Table 1
Epigenetics research contents and detection techniques"
| 研究内容 Research content | 高通量检测技术 High throughput detection technology |
| DNA甲基化DNA methylation | WGBS、BS-Seq、RRBS、MeDIP-seq等 |
| RNA修饰RNA modification | MeRIP-seq/m6A-seq、m6Aseq 2、msC-seq、m7G-seq、m6A-REF-seq、m6A-LAIC-seq、DART-Seq等 |
| 染色质可及性Chromatin accessibility | DNase-seq、MNase-seq、FAIRE-seq、ATAC-seq、scATAC-seq等 |
| 组蛋白修饰Histone modification | ChIP-seq、CUT&Tag、CUT&Run、DAP-seq、HiChIP等 |
| 三维基因组3D genomics | Hi-C、scHi-C、Micro-C等 |

Fig. 2
3D structure of the genome(revised from [28]) Chromatin is highly coiled and folded within the cell nucleus and presents an ordered organizational structure. Chromatin is functionally and structurally classified into Compartment A and Compartment B. Compartment A is mainly composed of euchromatin and represents the active gene transcription region, while Compartment B is mainly composed of heterochromatin and represents the silent gene transcription region. Compartment A usually contains TADs, which can mediate the interaction between distant genes. CTCF and Cohesin form Loop structures of chromatin within the cell nucleus"
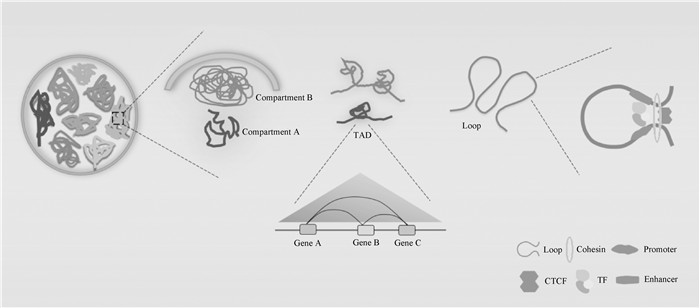
Table 2
Application of epigenetic regulation involving non-coding RNA in sheep genetic breeding"
| 物种 Species | 品种 Breed | 性状 Trait | 调控机制 Regulation mechanism | 非编码RNA ncRNA | 基因 Gene | 参考文献 Reference |
| 山羊 Goat | 陕北白绒山羊 | 绒毛性状 | DNA甲基化 | lnc_000374、lnc_002056 | HOXC13、SOX9、SOX21、JUNB、LHX2、VDR、GATA3 | [ |
| 安淮山羊 | 脂质代谢 | DNA甲基化 | XLOC_960044、XLOC_767346 | CACNA1E | [ | |
| 简州大耳羊 | 生长性状 | RNA修饰 | miR-503-5p | Ythdf2 | [ | |
| 简州大耳羊 | 生长性状 | RNA修饰 | miR-874-3p、miR-874-3p、circRNA_0873、circRNA_0873、circRNA_1161 | LAMA5、EBF3、HDAC11、CCND2 | [ | |
| 绒山羊 | 绒毛性状 | RNA修饰 | circPAPPA | PAPPA | [ | |
| 内蒙古绒山羊 | 绒毛性状 | RNA修饰 | circRNA_2130、circRNA_0013、circRNA_1203、circRNA_1462、circRNA_1242、circRNA_2308、circRNA_2654、circRNA_1442 | ANGEL2、APP、GKAP1、HNRNPC、PTBP3、NUCB1、SNRK、ZNF609 | [ | |
| 辽宁绒山羊 | 绒毛性状 | RNA修饰 | circHECA | HECA | [ | |
| 辽宁绒山羊 | 绒毛性状 | RNA修饰 | circERCC6 | ERCC6 | [ | |
| 辽宁绒山羊 | 绒毛性状 | RNA修饰 | circRNA-ZNF638 | ZNF638 | [ | |
| 黑山羊 | 繁殖性状 | RNA修饰 | circRNA4464、circRNA1212、circRNA1213、circRNA1149、circRNA4524等 | — | [ | |
| 绵羊 Sheep | 湖羊 | 生长性状 | DNA甲基化 | lncRNA | GTL2 | [ |
| 小尾寒羊 | 绒毛性状 | RNA修饰 | lncRNA: MSTRG.46299-PSEN2、ENSOARG00020016306-CCND3、ENSOARG00020002712-COL2A1、ENSOARG00020008516-ERCC3 | PSEN2、CCND3、COL2A1、ERCC3 | [ |
| 1 |
MISTELI T . The self-organizing genome: Principles of genome architecture and function[J]. Cell, 2020, 183 (1): 28- 45.
doi: 10.1016/j.cell.2020.09.014 |
| 2 |
MATTEI A L , BAILLY N , MEISSNER A . DNA methylation: a historical perspective[J]. Trends Genet, 2022, 38 (7): 676- 707.
doi: 10.1016/j.tig.2022.03.010 |
| 3 |
RUZOV A , TSENKINA Y , SERIO A , et al. Lineage-specific distribution of high levels of genomic 5-hydroxymethylcytosine in mammalian development[J]. Cell Res, 2011, 21 (9): 1332- 1342.
doi: 10.1038/cr.2011.113 |
| 4 |
ILLINGWORTH R S , GRUENEWALD-SCHNEIDER U , WEBB S , et al. Orphan CpG islands identify numerous conserved promoters in the mammalian genome[J]. PLoS Genet, 2010, 6 (9): e1001134.
doi: 10.1371/journal.pgen.1001134 |
| 5 | 谢银平, 肖玲, 郑雅格, 等. DNA甲基化在抑郁症研究中的进展[J]. 神经损伤与功能重建, 2022, 17 (5): 277- 280. |
| XIE Y P , XIAO L , ZHENG Y G , et al. Progress of DNA methylation in depression research[J]. Neural Injury and Functional Reconstruction, 2022, 17 (5): 277- 280. | |
| 6 |
URICH M A , NERY J R , LISTER R , et al. MethylC-seq library preparation for base-resolution whole-genome bisulfite sequencing[J]. Nat Protoc, 2015, 10 (3): 475- 483.
doi: 10.1038/nprot.2014.114 |
| 7 |
MEISSNER A , GNIRKE A , BELL G W , et al. Reduced representation bisulfite sequencing for comparative high-resolution DNA methylation analysis[J]. Nucleic Acids Res, 2005, 33 (18): 5868- 5877.
doi: 10.1093/nar/gki901 |
| 8 |
BOCCALETTO P , STEFANIAK F , RAY A , et al. MODOMICS: a database of RNA modification pathways[J]. Nucleic Acids Res, 2022, 50 (D1): D231- D235.
doi: 10.1093/nar/gkab1083 |
| 9 |
LIU Q , GREGORY R I . RNAmod: an integrated system for the annotation of mRNA modifications[J]. Nucleic Acids Res, 2019, 47 (W1): W548- w555.
doi: 10.1093/nar/gkz479 |
| 10 |
MEYER K D , SALETORE Y , ZUMBO P , et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3' UTRs and near stop codons[J]. Cell, 2012, 149 (7): 1635- 1646.
doi: 10.1016/j.cell.2012.05.003 |
| 11 |
XUE T X , QIU X Y , LIU H Y , et al. Epigenetic regulation in fibrosis progress[J]. Pharmacol Res, 2021, 173, 105910.
doi: 10.1016/j.phrs.2021.105910 |
| 12 | WANG Y , LI Y , YUE M , et al. N6-methyladenosine RNA modification regulates embryonic neural stem cell self-renewal through histone modifications[J]. Nat Neurosci, 2018, 21 (8): 1139. |
| 13 | WU J , QUAN J P , YE Y , et al. [Advances in assay for transposase-accessible chromatin with high-throughput sequencing][J]. Yi Chuan, 2020, 20: 42 (4): 333- 346. |
| 14 |
COCKERILL P N . Structure and function of active chromatin and DNase Ⅰ hypersensitive sites[J]. FEBS J, 2011, 278 (13): 2182- 2210.
doi: 10.1111/j.1742-4658.2011.08128.x |
| 15 | CHEN A , CHEN D Z , CHEN Y . Advances of DNase-seq for mapping active gene regulatory elements across the genome in animals[J]. Gene, 2018, 15: 667, 83- 94. |
| 16 | 欧阳也, 秦玉婷, 姚超, 等. 利用ATAC-seq技术在人免疫细胞中检测染色质开放性的方法建立[J]. 现代免疫学, 2020, 40 (2): 93- 99. |
| OUYANG Y , QIN Y T , YAO C , et al. Establishment of a method to detect chromatin openness in human immune cells using ATAC-seq[J]. Modern Immunology, 2020, 40 (2): 93- 99. | |
| 17 |
JAMBHEKAR A , DHALL A , SHI Y . Roles and regulation of histone methylation in animal development[J]. Nat Rev Mol Cell Biol, 2019, 20 (10): 625- 641.
doi: 10.1038/s41580-019-0151-1 |
| 18 |
LIU C F , ABOUSI A , BAZELEY P , et al. Global analysis of histone modifications and long-range chromatin interactions revealed the differential cistrome changes and novel transcriptional players in human dilated cardiomyopathy[J]. J Mol Cell Cardiol, 2020, 145, 30- 42.
doi: 10.1016/j.yjmcc.2020.06.001 |
| 19 |
MORGAN M A , SHILATIFARD A . Reevaluating the roles of histone-modifying enzymes and their associated chromatin modifications in transcriptional regulation[J]. Nat Genet, 2020, 52 (12): 1271- 1281.
doi: 10.1038/s41588-020-00736-4 |
| 20 | SUN P , HUANG T R , HUANG C , et al. Role of histone modification in the occurrence and development of osteoporosis[J]. Front Endocrinol (Lausanne), 2022, 26: 13, 964103. |
| 21 |
CHEN Z Y , DJEKIDEL M N , ZHANG Y . Distinct dynamics and functions of H2AK119ub1 and H3K27me3 in mouse preimplantation embryos[J]. Nat Genet, 2021, 53 (4): 551- 563.
doi: 10.1038/s41588-021-00821-2 |
| 22 |
WANG Y F , YANG L , ZHANG X J , et al. Epigenetic regulation of ferroptosis by H2B monoubiquitination and p53[J]. EMBO Rep, 2019, 20 (7): e47563.
doi: 10.15252/embr.201847563 |
| 23 | ALBERT I , MAVRICH T N , TOMSHO L P , et al. Translational and rotational settings of H2A. Z nucleosomes across the Saccharomyces cerevisiae genome[J]. Nature, 2007, 29:446 (7135): 572- 576. |
| 24 | SKENE P J , HENIKOFF S . An efficient targeted nuclease strategy for high-resolution mapping of DNA binding sites[J]. Elife, 2017, 16: 6, e21856. |
| 25 |
KAYA-OKUR H S , JANSSENS D H , HENIKOFF J G , et al. Efficient low-cost chromatin profiling with CUT&Tag[J]. Nat Protoc, 2020, 15 (10): 3264- 3283.
doi: 10.1038/s41596-020-0373-x |
| 26 | 邱格格, 林胜男, 黄浩. 三维基因组学及其应用概述[J]. 生物学教学, 2021, 46 (10): 4- 6. |
| QIU G G , LIN S N , HUANG H . Overview of three-dimensional genomics and its applications[J]. Teaching Biology, 2021, 46 (10): 4- 6. | |
| 27 | LUPIÁÑEZ D G , KRAFT K , HEINRICH V , et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions[J]. Cell, 2015, 21: 161 (5): 1012- 1025. |
| 28 |
MATHARU N , AHITVV N . Minor loops in major folds: Enhancer-promoter looping, chromatin restructuring, and their association with transcription regulation and disease[J]. PLoS Genet, 2015, 11 (12): e1005640.
doi: 10.1371/journal.pgen.1005640 |
| 29 | DEKKER J , RIPPE K , DEKKER M , et al. Capturing chromosome conformation[J]. Science, 2002, 15: 295 (5558): 1306- 1311. |
| 30 |
SIMONIS M , KlLOUS P , SPLINTER E , et al. Nuclear organization of active and inactive chromatin domains uncovered by chromosome conformation capture-on-chip (4C)[J]. Nat Genet, 2006, 38 (11): 1348- 1354.
doi: 10.1038/ng1896 |
| 31 |
DOSTIE J , RICHMOND T A , ARNAOUT R A , et al. Chromosome Conformation Capture Carbon Copy (5C): a massively parallel solution for mapping interactions between genomic elements[J]. Genome Res, 2006, 16 (10): 1299- 1309.
doi: 10.1101/gr.5571506 |
| 32 | LIEBERMAN-AIDEN E , VAN BERKUM N L , WILLIAMS L , et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome[J]. Science, 2009, 9: 326 (5950): 289- 293. |
| 33 | FAN Y , LIANG Y , DENG K , et al. Analysis of DNA methylation profiles during sheep skeletal muscle development using whole-genome bisulfite sequencing[J]. BMC Genomics, 2020, 29: 21 (1): 327. |
| 34 | 岳彩娟, 梁小军, 王秀琴, 等. 滩羊和湖羊背最长肌全基因组甲基化差异分析[J]. 中国畜牧兽医, 2020, 47 (10): 3058- 3068. |
| YUE C J , LIANG X J , WANG X Q , et al. Analysis of genome-wide methylation differences in the longest dorsal muscle of beach and lake sheep[J]. China Animal Husbandry and Veterinary Medicine, 2020, 47 (10): 3058- 3068. | |
| 35 | YUE C Y , WANG J K , SHEN Y F , et al. Whole-genome DNA methylation profiling reveals epigenetic signatures in developing muscle in Tan and Hu sheep and their offspring[J]. Front Vet Sci, 2023, 14: 10, 1186040. |
| 36 |
FAN Y X , REN C F , DENG K P , et al. The regulation of LncRNA GTL2 expression by DNA methylation during sheep skeletal muscle development[J]. Genomics, 2022, 114 (5): 110453.
doi: 10.1016/j.ygeno.2022.110453 |
| 37 | 罗建兴, 刘国强, 张宏博, 等. 蒙古羊后腿肌肉组织中MYF6和MEF2A基因的甲基化分析[J]. 黑龙江畜牧兽医, 2021 (3): 22- 26. |
| LUO J X , LIU G Q , ZHANG H B , et al. Methylation analysis of MYF6 and MEF2A genes in muscle tissues of hind legs of Mongolian sheep[J]. Heilongjiang Animal Husbandry and Veterinary Medicine, 2021 (3): 22- 26. | |
| 38 | 侯晨曦, 洪文娟, 何宗龙, 等. LEF1基因在巴什拜羊不同毛色皮肤组织中的DNA甲基化及mRNA表达水平分析[J]. 新疆农业科学, 2022, 59 (11): 2742- 2748. |
| HOU C X , HONG W J , HE Z L , et al. Analysis of DNA methylation and mRNA expression levels of LEF1 gene in skin tissues of Bashbay sheep with different coat colours[J]. Xinjiang Agricultural Science, 2022, 59 (11): 2742- 2748. | |
| 39 | XIAO P , ZHONG T , LIU Z F , et al. Integrated analysis of methylome and transcriptome changes reveals the underlying regulatory signatures driving curly wool transformation in Chinese Zhongwei goats[J]. Front Genet, 2020, 8: 10, 1263. |
| 40 | LI C , LI Y , ZHOU G X , et al. Whole-genome bisulfite sequencing of goat skins identifies signatures associated with hair cycling[J]. BMC Genomics, 2018, 28: 19 (1): 638. |
| 41 | WANG S H , LI F , LIU J W , et al. Integrative Analysis of Methylome and Transcriptome Reveals the Regulatory Mechanisms of Hair Follicle Morphogenesis in Cashmere Goat[J]. Cells, 2020, 14: 9 (4): 969. |
| 42 | ZHANG Y L , LI F Z , FENG X , et al. Genome-wide analysis of DNA Methylation profiles on sheep ovaries associated with prolificacy using whole-genome Bisulfite sequencing[J]. BMC Genomics, 2017, 2: 18 (1): 759. |
| 43 | YANG C M , HE J M , MAO J Y , et al. Genome-wide DNA methylation analysis and functional validation of litter size traits in Jining grey goats[J]. Genes (Basel), 2024, 12: 15 (3): 353. |
| 44 |
KANG B , WANG J , ZHANG H , et al. Genome-wide profile in DNA methylation in goat ovaries of two different litter size populations[J]. J Anim Physiol Anim Nutr (Berl), 2022, 106 (2): 239- 249.
doi: 10.1111/jpn.13600 |
| 45 |
ZHENG L M , ZHAI Y X , LI N , et al. Modification of Tet1 and histone methylation dynamics in dairy goat male germline stem cells[J]. Cell Prolif, 2016, 49 (2): 163- 172.
doi: 10.1111/cpr.12245 |
| 46 | ZHENG L , ZHAI Y X , LI N , et al. The modification of Tet1 in male germline stem cells and interact with PCNA, HDAC1 to promote their self-renewal and proliferation[J]. Sci Rep, 2016, 18: 6, 37414. |
| 47 | WU H , ZHOU W D , LIU H J , et al. Whole-genome methylation analysis reveals epigenetic variation between wild-type and nontransgenic cloned, ASMT transgenic cloned dairy goats generated by the somatic cell nuclear transfer[J]. J Anim Sci Biotechnol, 2022, 25: 13 (1): 145. |
| 48 |
YANG C , GAO X , YE J , et al. The interaction between DNA methylation and long non-coding RNA during the onset of puberty in goats[J]. Reprod Domest Anim, 2018, 53 (6): 1287- 1297.
doi: 10.1111/rda.13246 |
| 49 | 刘波, 芮雪, 方翟, 等. 多浪羊与卡拉库尔羊FSHR基因启动子区甲基化水平差异研究[J]. 中国畜牧杂志, 2021, 57 (11): 126- 130. |
| LIU B , RUI X , FANG Z , et al. Differences in methylation levels between the promoter regions of FSHR genes in Dolang and Karakul sheep[J]. Chinese Journal of Animal Husbandry, 2021, 57 (11): 126- 130. | |
| 50 | 胡靖玮, 杨凯捷, 乔利英, 等. 绵羊脂肪组织AGPAT2基因启动子区DNA甲基化及其表达研究[J]. 中国畜牧杂志, 2023, 59 (2): 92- 97. |
| HU J W , YANG K J , QIAO L Y , et al. DNA methylation in the promoter region of AGPAT2 gene and its expression in sheep adipose tissue[J]. Chinese Journal of Animal Husbandry, 2023, 59 (2): 92- 97. | |
| 51 | ZHU C Y , SONG S Z , LI M N , et al. Genome-wide DNA methylation analysis reveals different methylation patterns in Chinese indigenous sheep with different type of tail[J]. Front Vet Sci, 2023, 5: 10, 1125262. |
| 52 | LUO R S , DAI X L , ZHANG L , et al. Genome-wide DNA methylation patterns of muscle and tail-fat in dairy Meade sheep and Mongolian sheep[J]. Animals (Basel), 2022, 29: 12 (11): 1399. |
| 53 | JIANG J F , GUO L Y , HUANG X , et al. Regulatory role of N6-Methyladenosine on skeletal muscle development in Hu sheep[J]. Front Genet, 2024, 21: 15, 1449144. |
| 54 | DENG K P , LIU Z P , LI X D , et al. Ythdf2-mediated STK11 mRNA decay supports myogenesis by inhibiting the AMPK/mTOR pathway[J]. Int J Biol Macromol, 2024, 254 (Pt 1): 127614. |
| 55 | DENG K , FAN Y X , LIANG Y X , et al. FTO-mediated demethylation of GADD45B promotes myogenesis through the activation of p38 MAPK pathway[J]. Mol Ther Nucleic Acids, 2021, 24: 26, 34- 48. |
| 56 | YAO J Z , XU L , ZHAO Z H , et al. Fat mass- and obesity-associated protein (fto) promotes the proliferation of goat skeletal muscle satellite cells by stabilizing DAG1 mRNA in an IGF2BP1-related m6A manner[J]. Int J Mol Sci, 2024, 11: 25 (18): 9804. |
| 57 |
SU Y L , DENG K P , LIU Z P , et al. m6A modified pre-miR-503-5p contributes to myogenic differentiation through the activation of mTOR pathway[J]. Int J Biol Macromol, 2025, 294, 139517.
doi: 10.1016/j.ijbiomac.2025.139517 |
| 58 | ZHAO S , CAO J X , SUN Y J , et al. METTL3 Promotes the Differentiation of Goat Skeletal Muscle Satellite Cells by Regulating MEF2C mRNA Stability in a m6A-Dependent Manner[J]. Int J Mol Sci, 2023, 14: 24 (18): 14115. |
| 59 | DENG K P , LIU Z P , LU X D , et al. Targeted Demethylation of the TGFβ1 mRNA Promotes Myoblast Proliferation via Activating the SMAD2 Signaling Pathway[J]. Cells, 2023, 24: 12 (7): 1005. |
| 60 | ZOU J H , SHEN Y J , ZOU J W , et al. Transcriptome-Wide Study Revealed That N6-Methyladenosine Participates in Regulation Meat Production in Goats[J]. Foods, 2023, 9: 12 (6): 1159. |
| 61 | WANG J M , LI X , QUBI W Q , et al. The Important Role of m6A-Modified circRNAs in the Differentiation of Intramuscular Adipocytes in Goats Based on MeRIP Sequencing Analysis[J]. Int J Mol Sci, 2023, 2: 24 (5): 4817. |
| 62 |
MENG J Z , LI J P , ZHAO Y Y . Comprehensive analysis of lncRNAs modified by m6A methylation in sheep skin[J]. Anim Biosci, 2024, 37 (11): 1887- 1990.
doi: 10.5713/ab.24.0039 |
| 63 |
ZHAO Y Y , MENG J Z , SONG X C , et al. m6A mRNA Methylation Analysis Provides Novel Insights into Pigmentation in Sheep Skin[J]. Epigenetics, 2023, 18 (1): 2230662.
doi: 10.1080/15592294.2023.2230662 |
| 64 | BAI M , SHEN J C , FAN Y X , et al. N6-methyladenosine (m6A)-circular RNA Pappalysin 1 (circPAPPA) from cashmere goats: Identification, regulatory network and expression potentially regulated by methylation in secondary hair follicles within the first intron of its host gene[J]. Animals (Basel), 2025, 18: 15 (4): 581. |
| 65 | HUA G Y , YANG X , MA Y H , et al. m6A methylation analysis reveals networks and key genes underlying the coarse and fine wool traits in a full-sib merino family[J]. Biology (Basel), 2022, 9: 11 (11): 1637. |
| 66 | ZHANG R , LIANG J Y , LIU Z M , et al. MeRIP-seq data analysis and validation reveal the regulatory role of m6A modified circRNAs in the apoptosis of secondary hair follicle cells in Inner Mongolia cashmere goats[J]. Comp Biochem Physiol Part D Genom Proteom, 2025, 12: 54, 101419. |
| 67 |
SHEN J C , HUI T Y , BAI M , et al. N6-methyladenosine (m6A)-circHECA from secondary hair follicle of cashmere goats: identification, regulatory network and expression regulated potentially by methylation of its host gene promoter[J]. Anim Biosci, 2024, 37 (12): 2066- 2080.
doi: 10.5713/ab.24.0081 |
| 68 | ZHANG Q , FAN Y X , BAI M , et al. CircERCC6 positively regulates the induced activation of SHF stem cells in cashmere goats via the miR-412-3p/BNC2 axis in an m6A-dependent manner[J]. Animals (Basel), 2024, 5: 14 (2): 187. |
| 69 |
YIN R H , YIN R L , BAI M , et al. N6-Methyladenosine modification (m6A) of circRNA-ZNF638 contributes to the induced activation of SHF stem cells through miR-361-5p/Wnt5a axis in cashmere goats[J]. Anim Biosci, 2023, 36 (4): 555- 569.
doi: 10.5713/ab.22.0211 |
| 70 |
WANG Y R , LI G Q , ZHANG X J , et al. Analysis of m6A methylation in skin tissues of different sex Liaoning cashmere goats[J]. Anim Biotechnol, 2023, 34 (2): 310- 320.
doi: 10.1080/10495398.2021.1962897 |
| 71 | WANG Y , ZHANG Y Y , GUO D , et al. m6A methylation analysis of differentially expressed genes in skin tissues of coarse and fine type Liaoning cashmere goats[J]. Front Genet, 2020, 22: 10, 1318. |
| 72 | LI D X , ZHOU L , LIU Z F , et al. FTO demethylates regulates cell-cycle progression by controlling CCND1 expression in luteinizing goat granulosa cells[J]. Theriogenology, 2024, 1: 216, 20- 29. |
| 73 | SUN Y , ZHANG X C , LI M D , et al. METTL3 promotes proliferation of goat endometrial epithelial cells by regulating CTGF in an m6A-dependent manner†[J]. Biol Reprod, 2023, 9: 108 (6): 902- 911. |
| 74 |
LIU Z F , LI D X , DENG M T , et al. METTL3 improves the development of somatic cell nuclear transfer embryos through AURKB and H3S10ph in goats[J]. Int J Biol Macromol, 2025, 286, 138546.
doi: 10.1016/j.ijbiomac.2024.138546 |
| 75 | LIU J , GUO C H , FU J J , et al. Identification and Functional Analysis of circRNAs during Goat Follicular Development[J]. Int J Mol Sci, 2024, 9: 25 (14): 7548. |
| 76 | LI D X , LIU Z F , DENG M T , et al. The function of the m6A methyltransferase METTL3 in goat early embryo development under hypoxic and normoxic conditions[J]. Theriogenology, 2022, 1: 177, 140- 150. |
| 77 | XI B P , LU Z K , ZHANG R , et al. Comprehensive analysis of the transcriptome-wide m6A Methylome in sheep testicular development[J]. Genomics, 2025, 22, 111005. |
| 78 | NABI KHAN R I , PRAHARAG M R , MALLA W A , et al. Changes in m6A RNA methylation of goat lung following PPRV infection[J]. Heliyon, 2023, 22: 9 (9): e19358. |
| 79 | KHAN O , TANUJ G N , CHORAVADA D R , et al. N6-Methyladenosine RNA Modification in Host Cells Regulates Peste des Petits Ruminants Virus Replication[J]. Microbiol Spectr, 2023, 14: 11 (2): e0266622. |
| 80 | SONG B W , WANG X , LIANG Z M , et al. RMDisease V2.0: an updated database of genetic variants that affect RNA modifications with disease and trait implication[J]. Nucleic Acids Res, 2023, 6: 51 (D1): D1388- D1396. |
| 81 |
CHEN B W , YUAN C , GUO T T , et al. The molecular regulated mechanism of METTL3 and FTO in lipid metabolism of Hu sheep[J]. Genomics, 2024, 116 (6): 110945.
doi: 10.1016/j.ygeno.2024.110945 |
| 82 | CHEN B W , YUAN C , GUO T T , et al. METTL3 and FTO Regulate Heat Stress Response in Hu Sheep Through Lipid Metabolism via m6A Modification[J]. Animals (Basel), 2025, 13: 15 (2): 193. |
| 83 | LU Z K , LIU J B , YUAN C , et al. m6A mRNA methylation analysis provides novel insights into heat stress responses in the liver tissue of sheep[J]. Genomics, 2021, 113 (1 Pt 2): 484- 492. |
| 84 |
LU Z K , MA Y J , LI Q , et al. The role of N6-methyladenosine RNA methylation in the heat stress response of sheep (Ovis aries)[J]. Cell Stress Chaperones, 2019, 24 (2): 333- 342.
doi: 10.1007/s12192-018-00965-x |
| 85 | CAO Y T , AI Y , ZHANG X S , et al. Genome-wide epigenetic dynamics during postnatal skeletal muscle growth in Hu sheep[J]. Commun Biol, 2023, 23: 6 (1): 1077. |
| 86 |
SU Y X , HE S Q , CHEN Q , et al. Integrative ATAC-seq and RNA-seq analysis of myogenic differentiation of ovine skeletal muscle satellite cell[J]. Genomics, 2024, 116 (3): 110851.
doi: 10.1016/j.ygeno.2024.110851 |
| 87 | CHEN Q L , CHEN Z , ZHANG Z X , et al. Profiling chromatin accessibility responses in goat bronchial epithelial cells infected with pasteurella multocida[J]. Int J Mol Sci, 2023, 9: 24 (2): 1312. |
| 88 |
BYRNE K , MCWILLIAM S , VUOCOLO T , et al. Genomic architecture of histone 3 lysine 27 trimethylation during late ovine skeletal muscle development[J]. Anim Genet, 2014, 45 (3): 427- 438.
doi: 10.1111/age.12145 |
| 89 | ALHARBI A B , SCHMITZ U , BAILEY C G , et al. CTCF as a regulator of alternative splicing: new tricks for an old player[J]. Nucleic Acids Res, 2021, 20: 49 (14): 7825- 7838. |
| 90 | YUAN Z , GE L , ZHANG W , et al. Preliminary results about lamb meat tenderness based on the study of novel isoforms and alternative splicing regulation pathways using Iso-seq, RNA-seq and CTCF ChIP-seq data[J]. Foods, 2022, 7: 11 (8): 1068. |
| 91 | DENG M T , LIU Z F , CHEN B B , et al. Aberrant DNA and histone methylation during zygotic genome activation in goat cloned embryos[J]. Theriogenology, 2020, 148, 27- 36. |
| 92 | LIU Z H , LI M Y , SUN Y , et al. Epigenetic dynamics of H4K20me3 modification during oocyte maturation and early reprogramming of somatic cell nuclear transfer goat embryos[J]. Am J Transl Res, 2022, 25: 14 (8): 5941- 5951. |
| 93 | MAO T C , HAN C Q , DENG R Z , et al. Treating donor cells with 2-PCPA corrects aberrant histone H3K4 dimethylation and improves cloned goat embryo development[J]. Syst Biol Reprod Med, 2018, 64 (3): 174- 182. |
| 94 | SINHA N , ROY S , HUANG B , et al. Developmental programming: prenatal testosterone-induced epigenetic modulation and its effect on gene expression in sheep ovary†[J]. Biol Reprod, 2020, 24: 102 (5): 1045- 1054. |
| 95 | BROOKS K E , BURNS G W , SPENCER T E . Peroxisome proliferator activator receptor gamma (PPARG) regulates conceptus elongation in sheep[J]. Biol Reprod, 2015, 92 (2): 42. |
| 96 | LI L Y , ZHANG D D , REN Y Y , et al. The modification of mitochondrial energy metabolism and histone of goat somatic cells under small molecules compounds induction[J]. Reprod Domest Anim, 2019, 54 (2): 138- 149. |
| 97 | GAO L , ZHANG Z H , ZHENG X M , et al. The novel role of Zfp296 in mammalian embryonic genome activation as an H3K9me3 modulator[J]. Int J Mol Sci, 2023, 12: 24 (14): 11377. |
| 98 | CHEN M , LONG X , CHEN M , et al. Integration of single-cell transcriptome and chromatin accessibility of early gonads development among goats, pigs, macaques, and humans[J]. Cell Rep, 2022, 1: 41 (5): 111587. |
| 99 | DAVENPORT K M , MASSA A T , BHATTARAI S , et al. Ovine FAANG project consortium. Characterizing genetic regulatory elements in ovine tissues[J]. Front Genet, 2021, 20: 12, 628849. |
| 100 | BILBAO-ARRIBAS M , JUGO B M . Transcriptomic meta-analysis reveals unannotated long non-coding RNAs related to the immune response in sheep[J]. Front Genet, 2022, 22: 13, 1067350. |
| 101 | MASSA A T , MOUSEL M R , DURFEE C J , et al. A DNA regulatory element haplotype at zinc finger genes is associated with host resilience to small ruminant lentivirus in two sheep populations[J]. Animals (Basel), 2021, 26: 11 (7): 1907. |
| 102 | WU J , LUO J , HE Q Y , et al. Docosahexaenoic acid alters lipid metabolism processes via H3K9ac epigenetic modification in dairy goat[J]. J Agric Food Chem, 2023, 71 (22): 8527- 8539. |
| 103 | HE Q Y , GAO L J H , ZHANG F H , et al. The FoxO1-ATGL axis alters milk lipolysis homeostasis through PI3K/AKT signaling pathway in dairy goat mammary epithelial cells[J]. J Anim Sci, 2023, 3: 101, 286. |
| 104 | WANG X , ZHANG F X , WANG Z M , et al. Histone H3K9 acetylation influences growth characteristics of goat adipose-derived stem cells in vitro[J]. Genet Mol Res, 2016, 15 (4): gmr15048954. |
| 105 | LI R , YANG P , DAI X L , et al. A near complete genome for goat genetic and genomic research[J]. Genet Sel Evol, 2021, 10: 53 (1): 74. |
| 106 | LI R , YANG P , LI M , et al. A Hu sheep genome with the first ovine Y chromosome reveal introgression history after sheep domestication[J]. Sci China Life Sci, 2021, 64 (7): 1116- 1130. |
| 107 | QIAO G Y , XU P , GUO T T , et al. Genetic basis of Dorper sheep (Ovis aries) revealed by long-read de novo genome assembly[J]. Front Genet, 2022, 11: 13, 846449. |
| 108 | BICKHART D M , ROSEN B D , KOREN S , et al. Single-molecule sequencing and chromatin conformation capture enable de novo reference assembly of the domestic goat genome[J]. Nat Genet, 2017, 49 (4): 643- 650. |
| 109 | DAVENPORT K M , BICKHART D M , WORLEY K , et al. An improved ovine reference genome assembly to facilitate in-depth functional annotation of the sheep genome[J]. Gigascience, 2022, 4: 11, 096. |
| 110 | GHURYE J , POP M , KOREN S , et al. Scaffolding of long read assemblies using long range contact information[J]. BMC Genomics, 2017, 12: 18 (1): 527. |
| 111 | WANG Z Y , LV Q , LI W Z , et al. Chromosome-level genome assembly of the cashmere goat[J]. Sci Data, 2024, 11 (1): 1107. |
| 112 | WU H , LUO L Y , ZHANG Y H , et al. Telomere-to-telomere genome assembly of a male goat reveals variants associated with cashmere traits[J]. Nat Commun, 2024, 15 (1): 10041. |
| [1] | ZHONG Xin, ZHANG Hui, ZHANG Chong, LIU Xiaohong. Research Progress on Genetic Breeding of Reproductive Performance in Sows [J]. Acta Veterinaria et Zootechnica Sinica, 2024, 55(2): 438-450. |
| [2] | RU Meng, ZENG Wenhui, PENG Jianling, ZENG Qingjie, YIN Chao, CUI Yong, WEI Qing, LIANG Haiping, XIE Xianhua, HUANG Jianzhen. Research Progress on Follicles Development of Hens and Its Epigenetic Regulatory Mechanism [J]. Acta Veterinaria et Zootechnica Sinica, 2023, 54(9): 3613-3622. |
| [3] | JIN Meilin, LI Taotao, SUN Dongxiao, WEI Caihong. Research Progress of Epigenetic Regulation in Fat Deposition Mechanism of Livestock and Poultry [J]. Acta Veterinaria et Zootechnica Sinica, 2023, 54(3): 855-867. |
| [4] | YANG Xiaogeng, ZHANG Huizhu, LI Jian, XIANG Hua, HE Honghong. Research Progress of the DNA Methylation in Mammalian Oocyte and Early Embryo Development [J]. Acta Veterinaria et Zootechnica Sinica, 2023, 54(2): 443-450. |
| [5] | LIU Yue, XUE Xianglan, LI Xiaobo, JIANG Lin, PU Yabin, HE Xiaohong, MA Yuehui, ZHAO Qianjun. Research Progress of the Relationship between Chromatin Accessibility and Animal Embryo Development [J]. Acta Veterinaria et Zootechnica Sinica, 2022, 53(3): 680-687. |
| [6] | WANG Di, YU Ying. Research Progress on Transcriptomics and Epigenetics of Bovine S. aureus Mastitis Resistance [J]. Acta Veterinaria et Zootechnica Sinica, 2022, 53(2): 329-338. |
| [7] | LU Zengkui, ZHANG Liping, LI Qing, LIU Enmin, DU Lixin, CHU Mingxing, WEI Caihong. N6-methyladenosine Modification in mRNA and Its Research Advance in Animals [J]. ACTA VETERINARIA ET ZOOTECHNICA SINICA, 2019, 50(1): 1-13. |
| [8] | ZHANG Tong-yu, WEI Xia, ZHANG Qin, DU Li-xin, WANG Li-xian, ZHAO Fu-ping. Progress on Application of Genomic Selection in Sheep and Goat Breeding [J]. ACTA VETERINARIA ET ZOOTECHNICA SINICA, 2018, 49(12): 2535-2542. |
| [9] | LIU Yan-li,SHEN Jing,ZHI Li-hui,LI Shi-zhao,YAO Jun-hu,YANG Xiao-jun. The Study on Epigenetic Mechanism of IGF2 Expression in Spleen and Thymus Regulated by Folic Acid in Broilers [J]. ACTA VETERINARIA ET ZOOTECHNICA SINICA, 2016, 47(2): 296-304. |
| [10] | WANG Kang-yan,LING Ying-hui,ZHANG Xiao-rong. Research Progress in Regulation of Gene Expression of Long Noncoding RNA [J]. ACTA VETERINARIA ET ZOOTECHNICA SINICA, 2015, 46(4): 509-517. |
| [11] | WANG Bo,DIAO Qi-yu. Advances on DNA Methylation and the Regulatory Effect of Nutrients on It [J]. ACTA VETERINARIA ET ZOOTECHNICA SINICA, 2015, 46(3): 349-356. |
| [12] | FU Yan-feng,LI Lan,ZHOU Yan-hong,LI Bi-xia,ZHAO Wei-min,ZHAO Fang,REN Shou-wen. Detection of EphA1 Expression and Methylation during Embryo Implantation in Meishan Pigs [J]. ACTA VETERINARIA ET ZOOTECHNICA SINICA, 2014, 45(12): 1964-1970. |
| [13] | KANG Qian,CUI Huan-zhong,SUN Yu-cheng,GAO Yun-hang,FAN Yi-wen,LIU Er-zhan. Research Progress on Regulation of Imprinted Genes for Animal Placental Phenotype [J]. ACTA VETERINARIA ET ZOOTECHNICA SINICA, 2013, 44(2): 169-173. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||